One quick way to get your data into Python is by using the Matlab API for Python.
First, install the MATLAB engine API for Python by following he instructions here. You will need to install as administrator.
Then, from your Python code you can simply do, as an example:
import matlab.engine
import numpy as np
from matplotlib import pyplot as plt
eng = matlab.engine.start_matlab()
z = eng.sbxread('d:\gm7\gm7_005_000',1,50)
q = np.array(z._data).reshape(z.size[::-1]).T
m = np.squeeze(np.ndarray.mean(q,axis=3))
plt.imshow(m,interpolation='nearest',cmap='gray')
plt.show()
This will read 50 frames starting at index 0, make it into a numpy array, take the mean over time, and display the result.
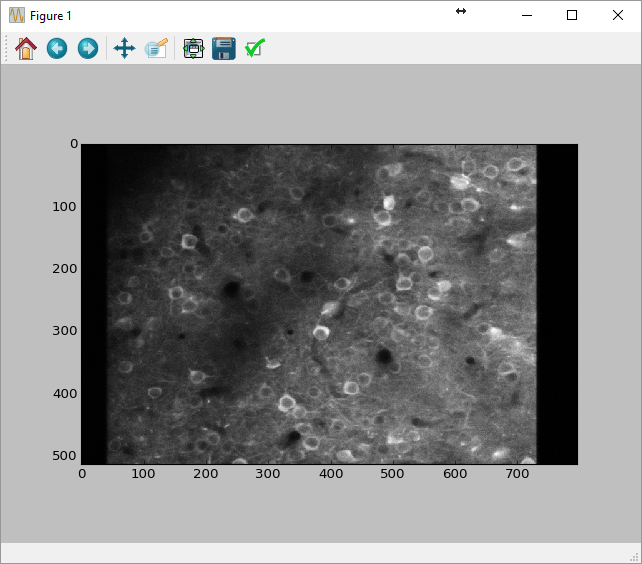
Creating the communication to the Matlab engine may take a few seconds, but after that the calls to sbxread() should be relatively fast. This is, of course, not the most efficient way to read the data into Python.
An alternative, offered by Francisco Luongo at Caltech, is to use the following native Python function:
import os
import scipy.io as spio
def loadmat(filename):
'''
this function should be called instead of direct spio.loadmat
as it cures the problem of not properly recovering python dictionaries
from mat files. It calls the function check keys to cure all entries
which are still mat-objects
'''
data = spio.loadmat(filename, struct_as_record=False, squeeze_me=True)
return _check_keys(data)
def _check_keys(dict):
'''
checks if entries in dictionary are mat-objects. If yes
todict is called to change them to nested dictionaries
'''
for key in dict:
if isinstance(dict[key], spio.matlab.mio5_params.mat_struct):
dict[key] = _todict(dict[key])
return dict
def _todict(matobj):
'''
A recursive function which constructs from matobjects nested dictionaries
'''
dict = {}
for strg in matobj._fieldnames:
elem = matobj.__dict__[strg]
if isinstance(elem, spio.matlab.mio5_params.mat_struct):
dict[strg] = _todict(elem)
else:
dict[strg] = elem
return dict
def sbxread(filename):
'''
Input: filename should be full path excluding .sbx
'''
# Check if contains .sbx and if so just truncate
if '.sbx' in filename:
filename = filename[:-4]
# Load info
info = loadmat(filename + '.mat')['info']
#print info.keys()
# Defining number of channels/size factor
if info['channels'] == 1:
info['nChan'] = 2; factor = 1
elif info['channels'] == 2:
info['nChan'] = 1; factor = 2
elif info['channels'] == 3:
info['nChan'] = 1; factor = 2
# Determine number of frames in whole file
max_idx = os.path.getsize(filename + '.sbx')/info['recordsPerBuffer']/info['sz'][1]*factor/4-1
# Paramters
k = 0; #First frame
N = max_idx; #Last frame
nSamples = info['sz'][1] * info['recordsPerBuffer'] * 2 * info['nChan']
# Open File
fo = open(filename + '.sbx')
# Note: There is a weird inversion that happns thus I am dding the negative sign....
fo.seek(k*nSamples, 0)
x = np.fromfile(fo, dtype = 'uint16',count = nSamples/2*N)
x = -x.reshape((info['nChan'], info['sz'][1], info['recordsPerBuffer'], N), order = 'F')
return x
Just a note that if you are trying to use Francisco’s code posted above on Window’s machine, the following line:
# Open File
fo = open(filename + ‘.sbx’)
needs to be changed to :
# Open File
fo = open(filename + ‘.sbx’,’rb’)
Thanks Mark… I still owe you a reply regarding network commands. Soon.